
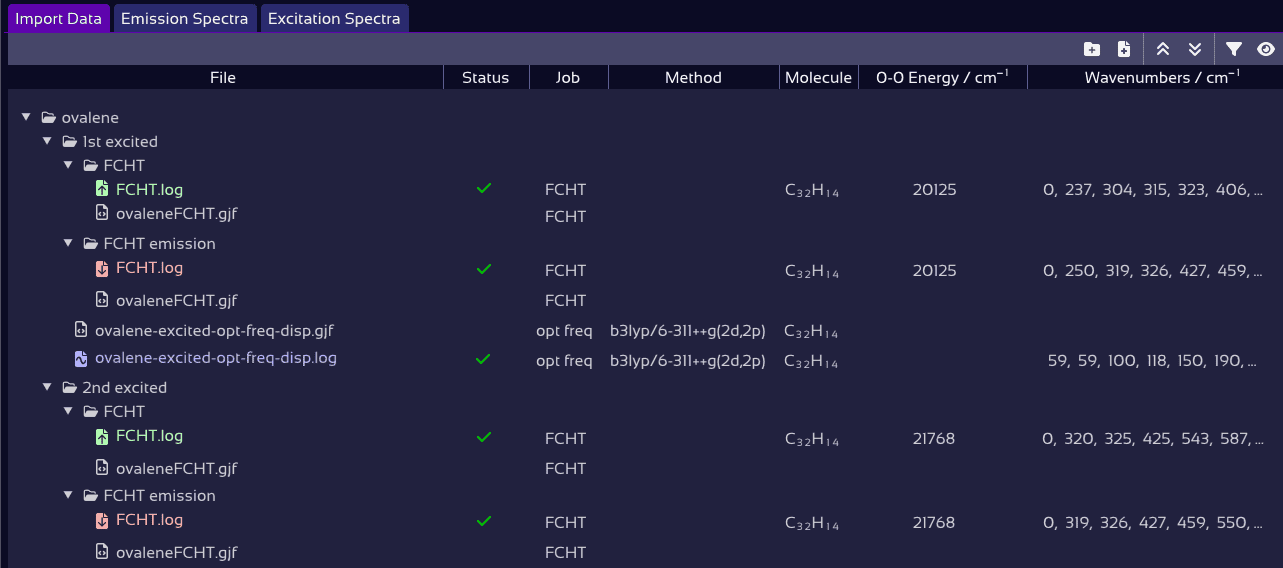
Gaussian jobs are scanned and labeled automatically during import.
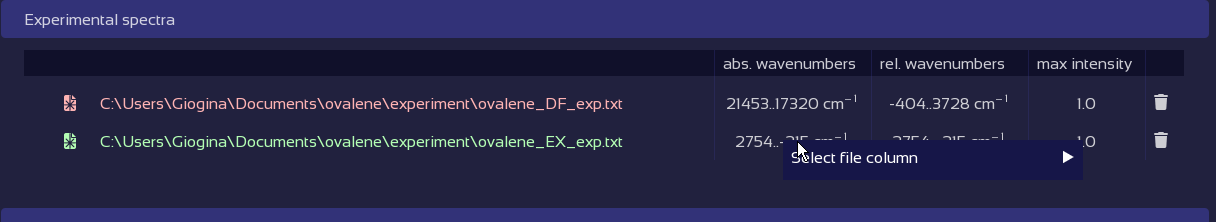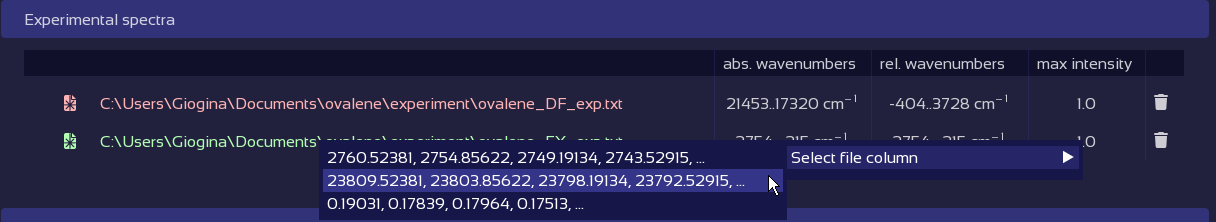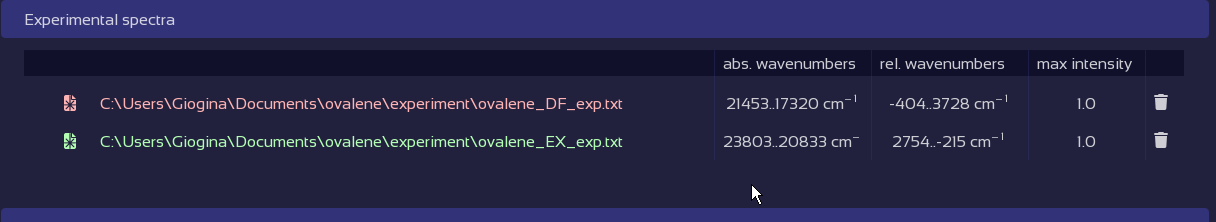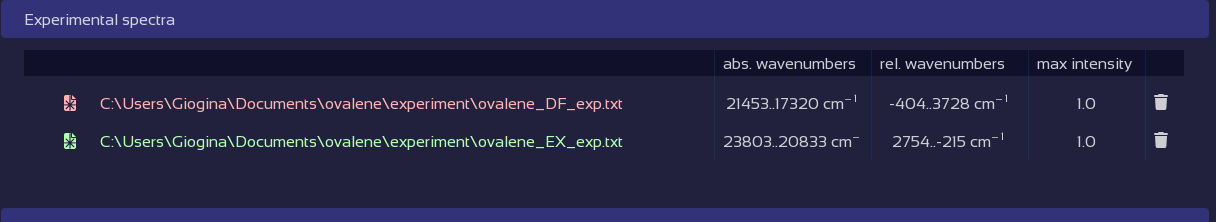
Fix broken data tables with a click.

Quickly shift wavenumbers or adjust peak broadening.
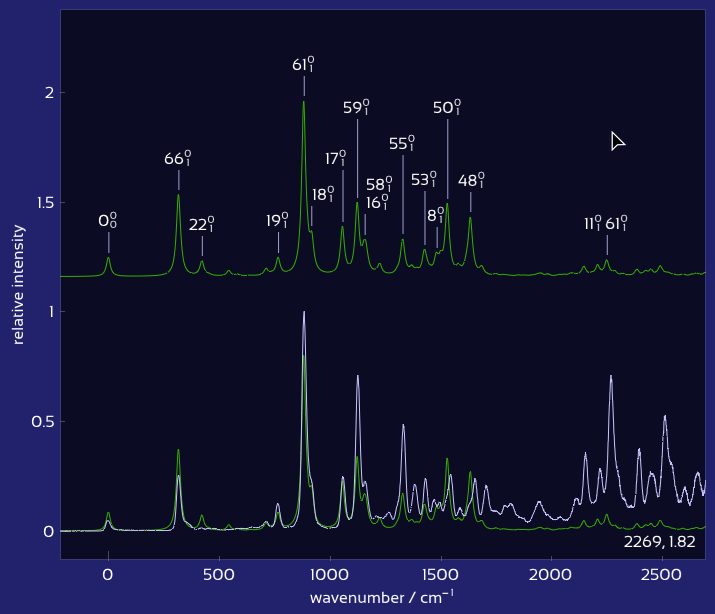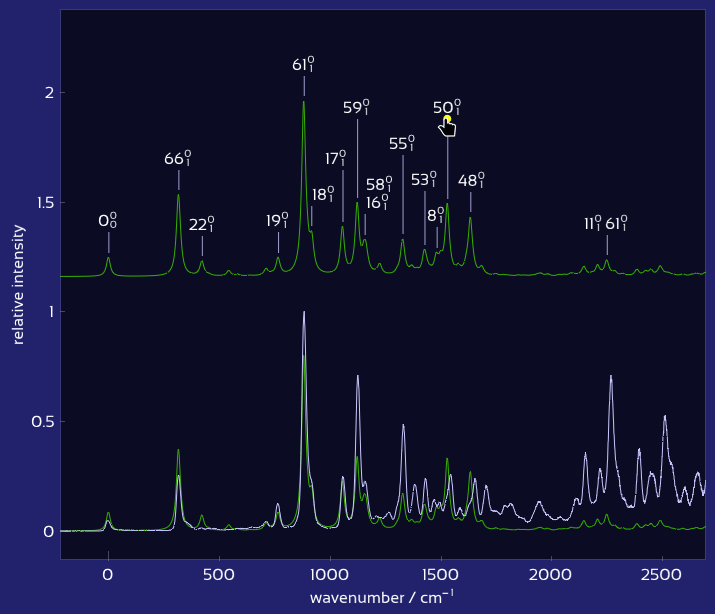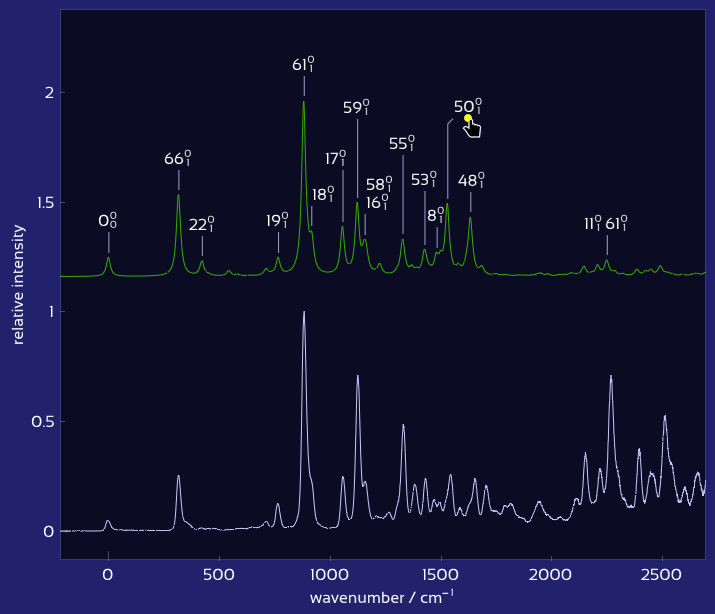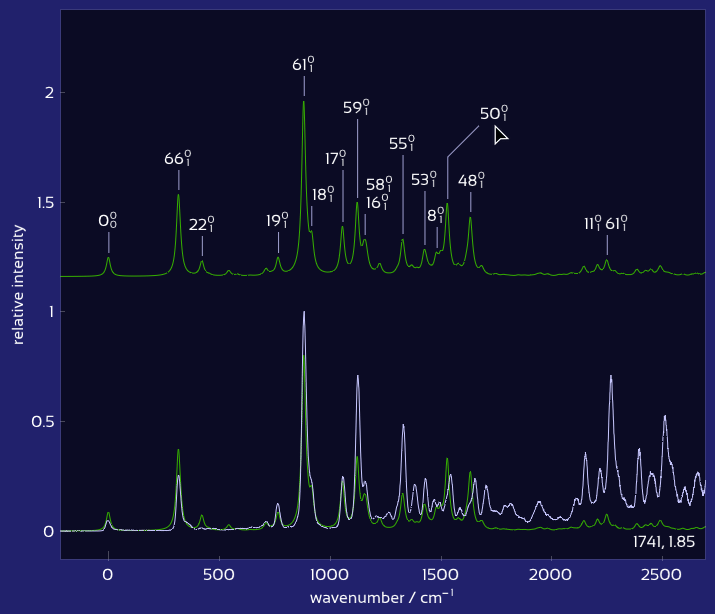
Drag labels to reposition them exactly where you want.
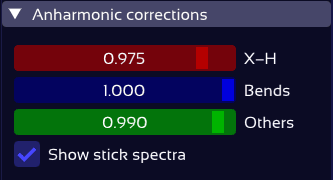

Each vibrational mode type gets its own scaling factor.
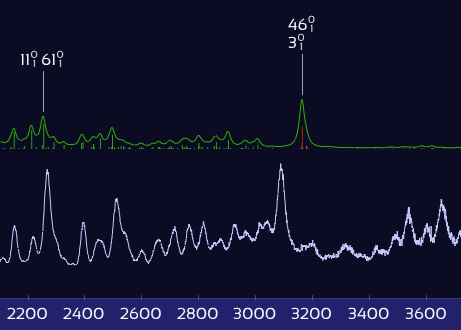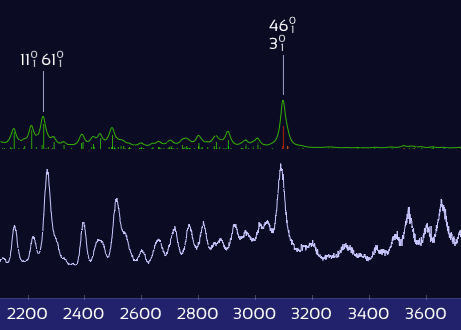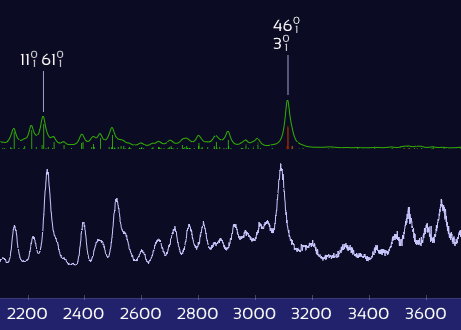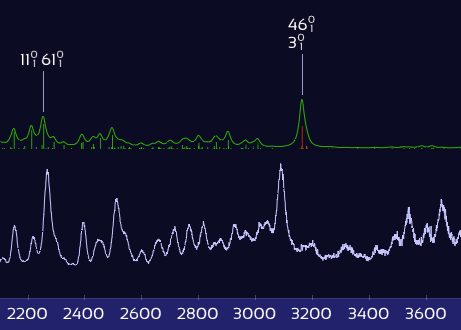
Only X–H stretch peaks shift — the rest stay put.
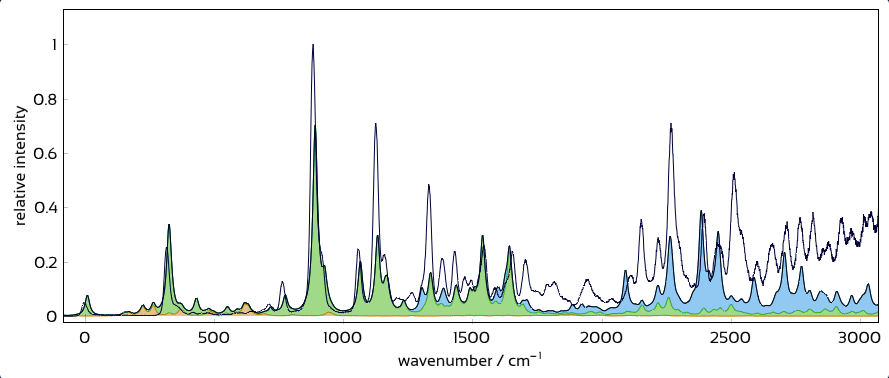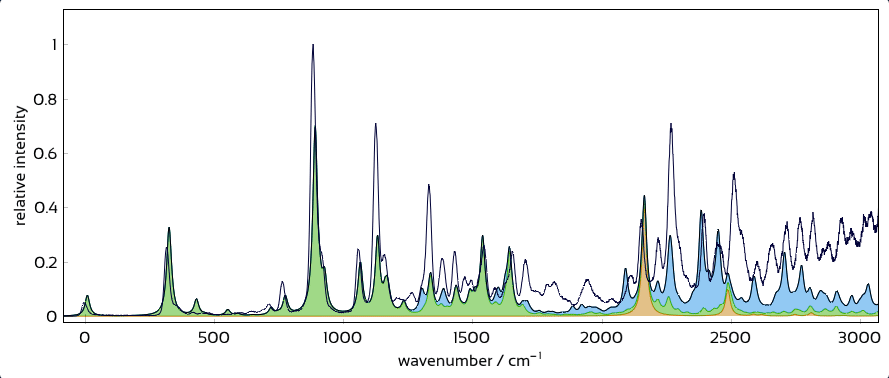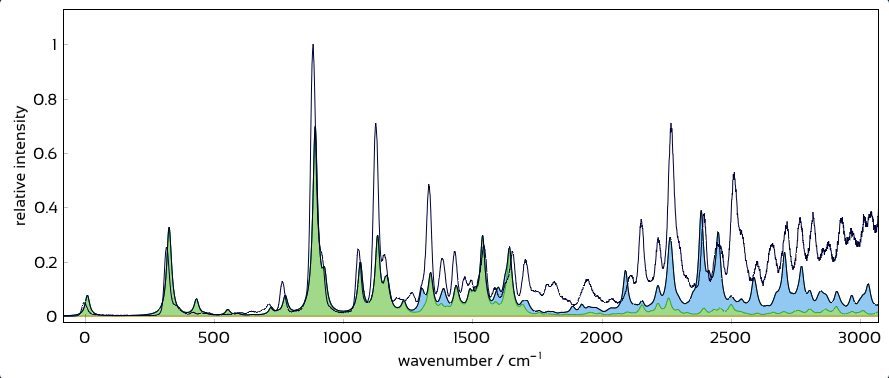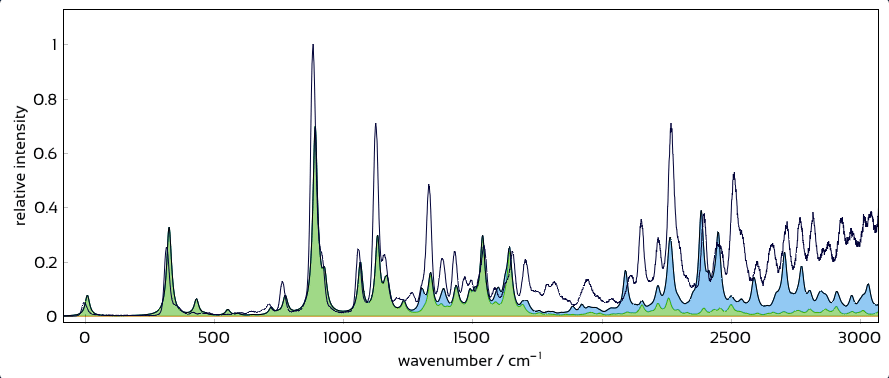
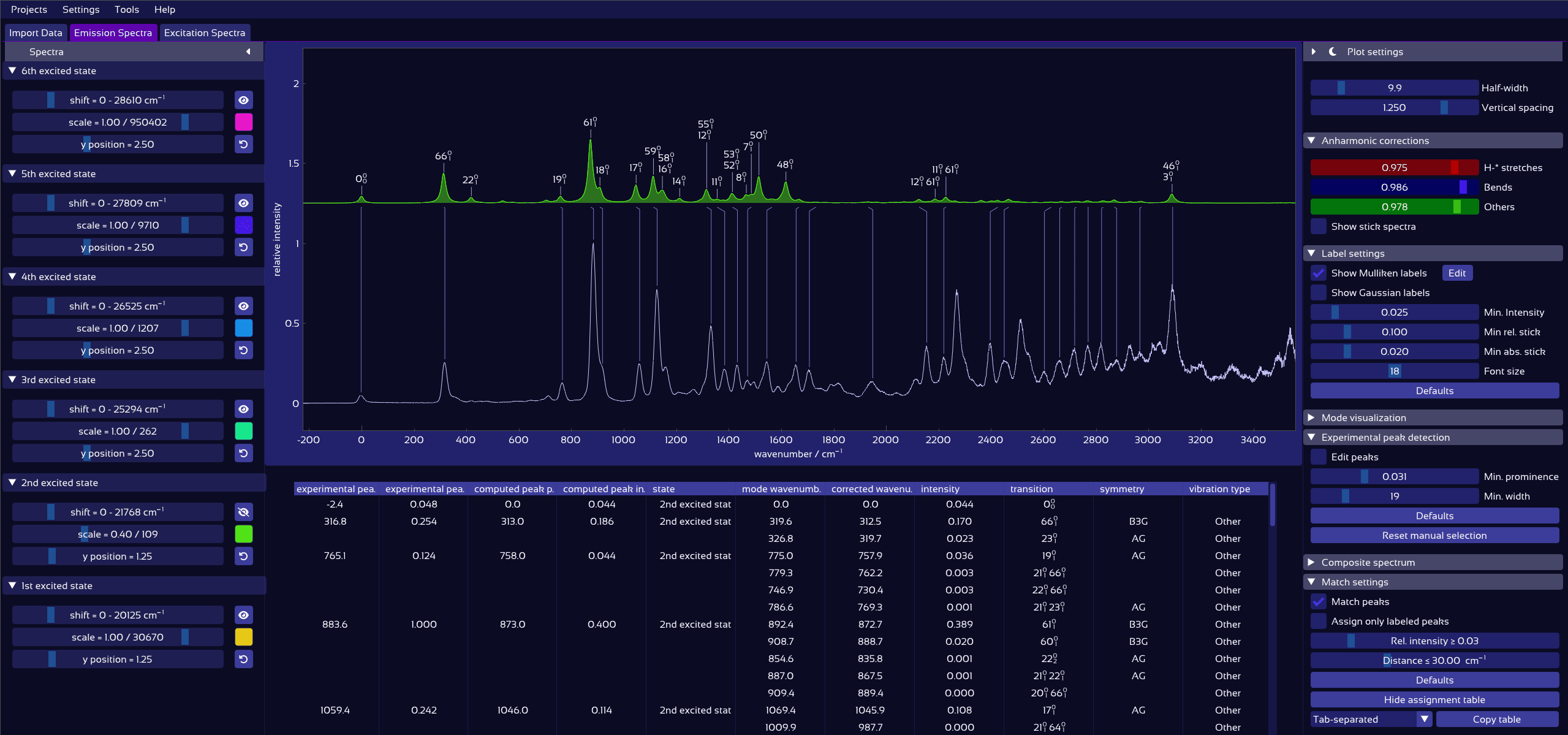
Interactively overlay excited-state contributions and see how well they explain experiment.
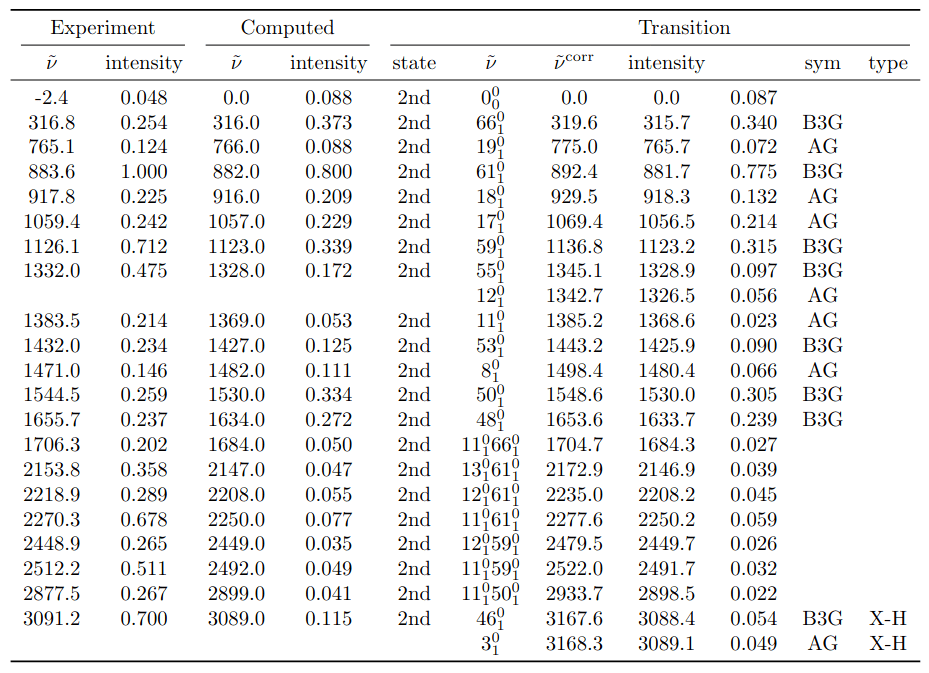
LaTeX export — no manual formatting required.
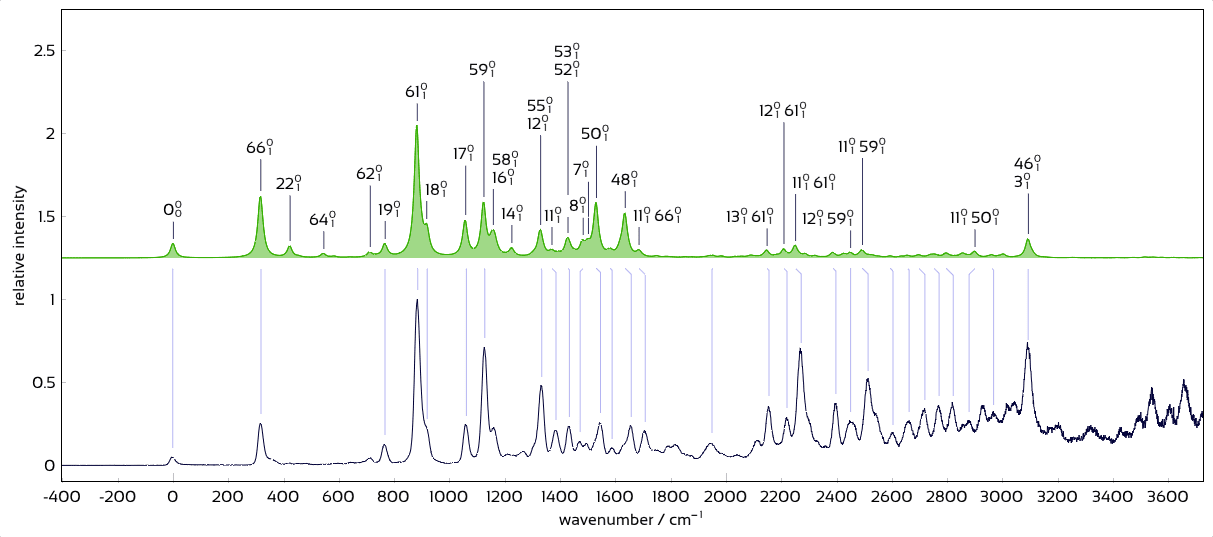
Plot of labeled, matched vibronic spectra.